Constraining Cosmological Parameters with Type Ia Supernovae Part I
In this problem, we will use data from Type Ia supernovae to constrain cosmological parameters.
This problem is based on The Supernova Cosmology Project, which is one of the two research teams that first determined a positive cosmological constant using redshift data from Type Ia supernovae. The team took data taken from various telescopes, including the HST to measure the magnitude of various supernovae as a function of their redshift. This work led to the 1997 discovery of the fact that the expansion of the universe is actually accelerating. The same work eventually won the 2011 Nobel Prize and the 2015 Breakthrough Prize in Fundamental Physics.
We use data from the said project for our analysis. The dataset we will use is called the SCP Union2.1 compilation. The compilation has data from 833 supernovae, drawn from 19 datasets but only a certain number pass usability cuts so we are going to use a subset of those 833 supernovae.
You can download the dataset from http://supernova.lbl.gov/Union/. Download the dataset that says Union2.1 Compilation Magnitude vs. Redshift Table (for your own cosmology fitter). Study the dataset and answer the following question.- How many usable supernovae are in the SCP Union2.1 dataset?
The columns are: supernova name, redshift, distance modulus, distance modulus error, and the probability that the supernova was hosted by a low-mass galaxy. Using this data, we can make what's called a Hubble diagram, which is a plot of distance against redshift.
Make the Hubble diagram with the given dataset. Your plot should look like the following.
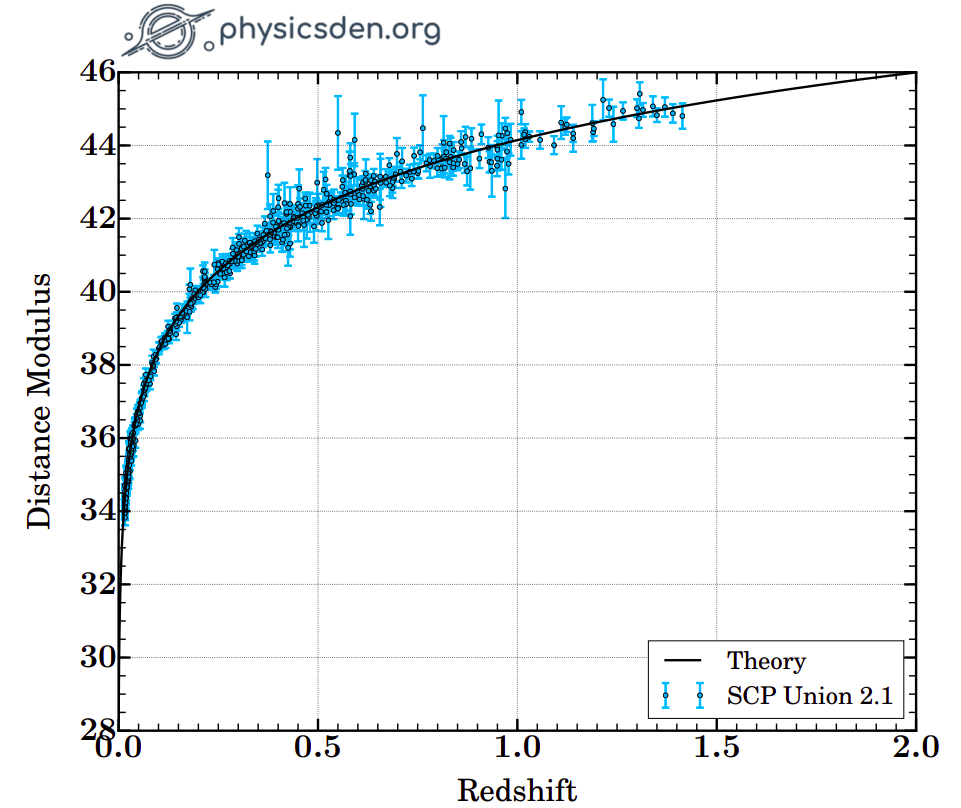
We can now write a Python function for our likelihood.
def calculate_likelihood(chi2):
# Implement the function here.
return likelihood
We're going to need the theoretical values of magnitude $m_{\mathrm{th}}$ so let's implement it but before that we need to implement the luminosity distance $d_L$. The luminosity distance depends on the comoving distance $d_{\mathrm{comov.}}$, which would in turn depend on the redshift $z$, equation of state $w$, and density parameters $\Omega_M$ and $\Omega_{\lambda}$. Implement this function and answer the questions below to check your implementation.
c = 299792 # Speed of light in km/s
H0 = 70 # Hubble constant
M = 25 # Nuisance parameter
def comoving_distance(z, w=-1, Omega_M=1.0, Omega_L=1.0):
# Implement the function here.
def luminosity_distance(z, w, Omega_M, Omega_L):
# Implement the function here.
return d_L
def m_th(z, w, Omega_M, Omega_L):
# Implement the function here.
# Calculate d_L with the function luminosity_distance() above.
return likelihood
- What is
comoving_distance(z=0.5, w=-1, Omega_M=0.25, Omega_L=0.75)? Answer to 1 decimal place.
- What is
luminosity_distance(z=0.5, w=-1, Omega_M=0.25, Omega_L=0.75)? Answer to 1 decimal place.
- What is
m_th(z=0.5, w=-1, Omega_M=0.25, Omega_L=0.75)? Answer to 1 decimal place.
For this problem, we will ignore the covariance matrix and assume it to be the identity matrix. The Union2.1 website does provide the covariance matrix if you want to use it to go beyond this problem.
We now have all the ingredients we need. We have the experimental measures of $m$ that we can calculate with the downloaded luminosity distances. We can now also calculate the theoretical values of $m$. With these two sets of information, we can vary our parameter values while looking at how that affects the likelihood. We want to maximize the likelihood, which would be equivalent to minimizing the $\chi^2$ value.
For this optimization, we need to calculate the $\chi^2$ values, so let's write a function to do so. Complete the code with hints given below. This should let you find the $\Omega_M$ value that maximizes the likelihood. You could also plot the likelihood as a function of $\Omega_M$. Your result should look like the image below.
def calculate_chi2(data, params):
z_s, mag_s, errors = data
omega_M, omega_DE, w = params
# Calculate chi2.
return chi2
def search_for_Omega_m():
z_s, mag_s, errors = load_data("SCPUnion2.1_mu_vs_z.txt")
w = -1
omega_M_s = np.linspace(0, 1, 1000)
# Calculate likelihoods.
# Find Omega_M that gives maximum likelihood.
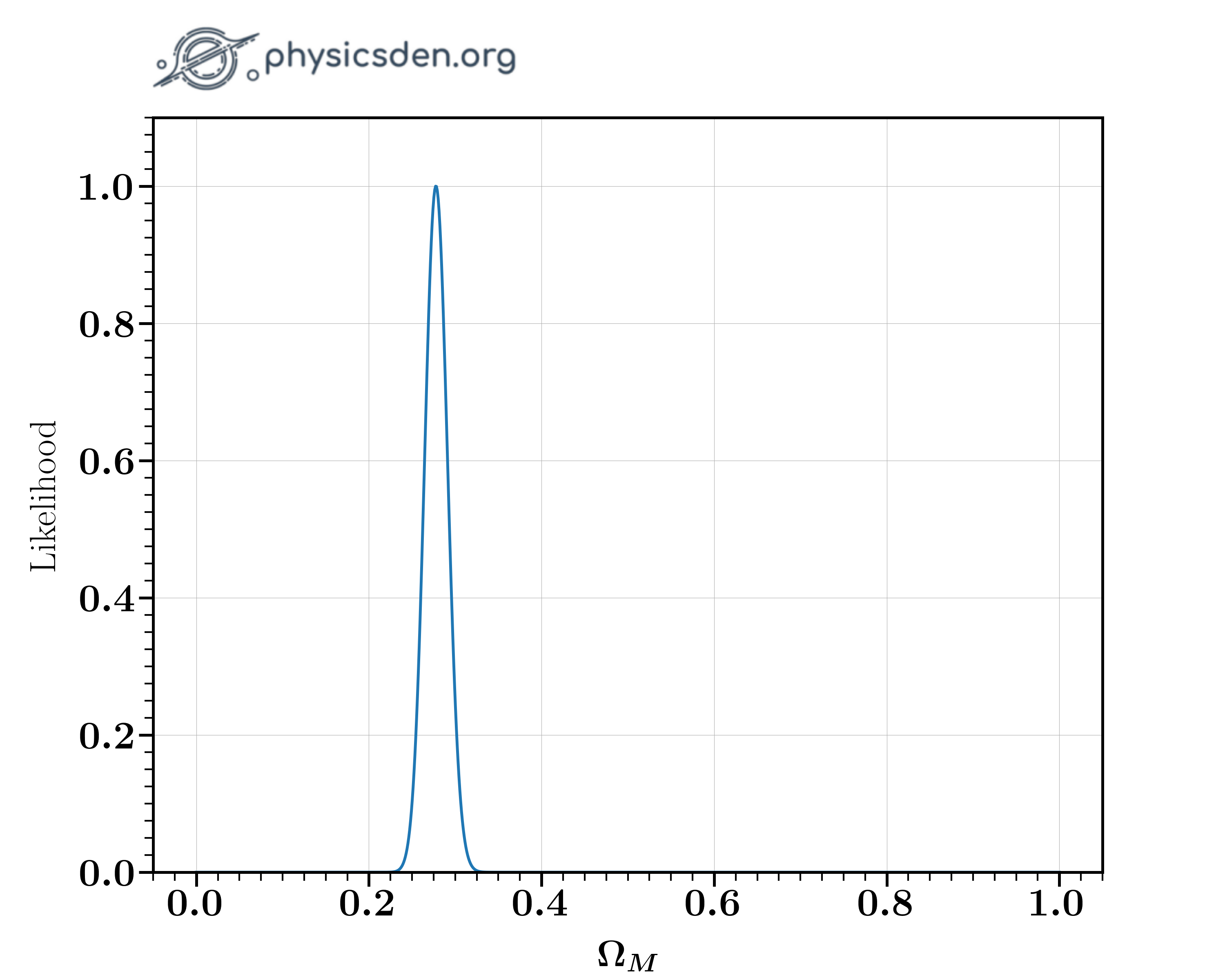
- What $\Omega_M$ gives you the maximum likelihood? Answer to 1 decimal place.
Now that we found the most likely value of $\Omega_M$, let's do a similar maximization to find constraints on two parameters simultaneously. In particular, let's optimize $\Omega_M, w$, and $\Omega_M, \Omega_{\Lambda}$.
Create a 2d grid of prospective parameter values and use that to maximize the likelihood. Fill in the code below to find the most likely values for $\Omega_M$ and $w$.
def search_for_Omega_m_and_w():
z_s, mag_s, errors = load_data("SCPUnion2.1_mu_vs_z.txt")
N = 50
omega_M_s = np.linspace(0, 0.5, N)
w_s = np.linspace(-1.5, 0, N)
likelihood_s = np.zeros((N, N))
# Loop through omega_M and w_s and find the optimum likelihood.
If you make a 2D contour plot of your likelihood, it should look something like the images below.
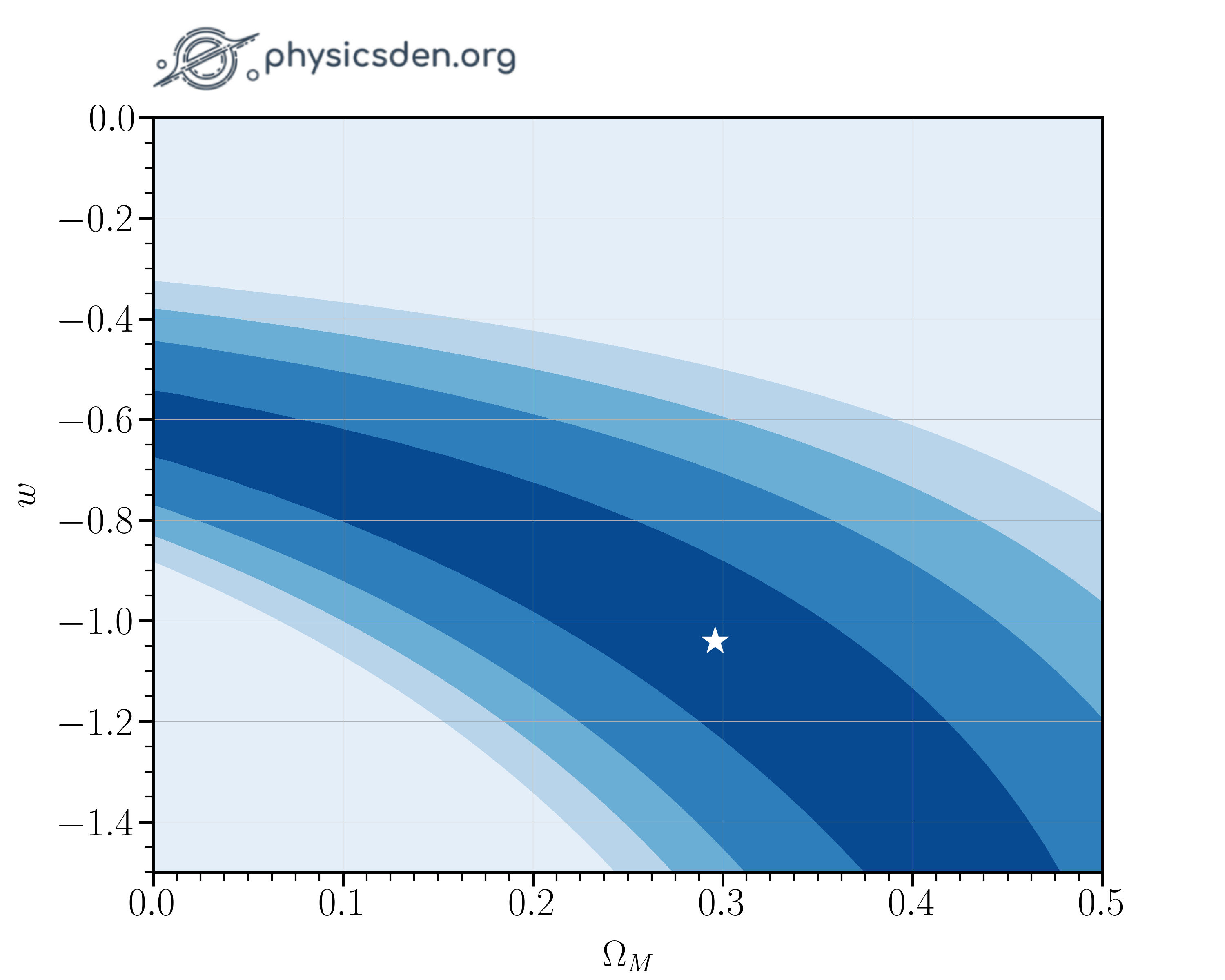
- What is the optimum $\Omega_M$? Answer to 1 decimal place.
- What is the optimum $w$? Answer to 1 decimal place.
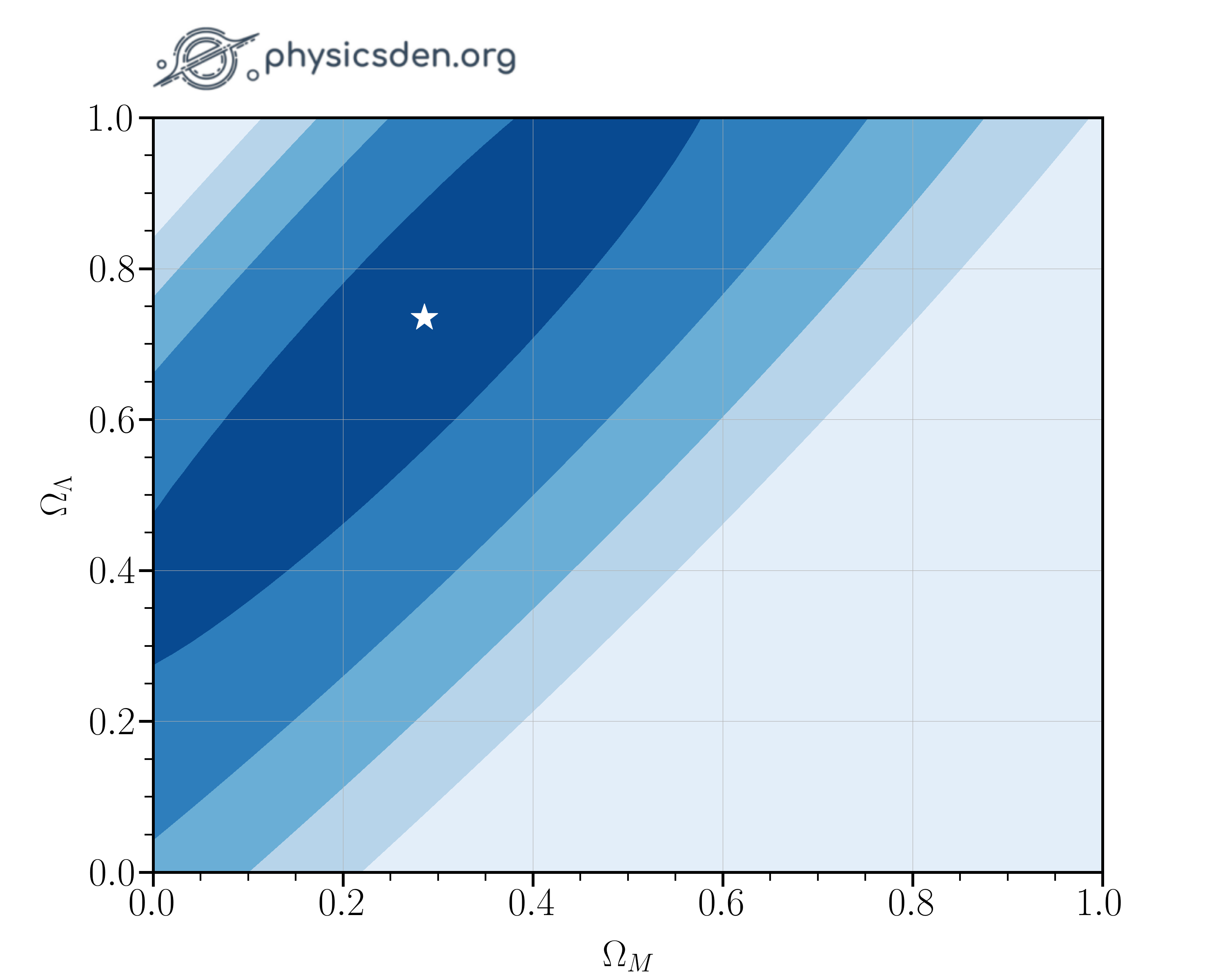
Repeat the steps you just took to find the most likely values for $\Omega_M$ and $\Omega_{\Lambda}$ simultaneously.
def search_for_Omega_m_and_Omega_L():
N = 50
omega_L_s = np.linspace(0.0, 1.0, N)
omega_M_s = np.linspace(0.0, 1.0, N)
likelihood_s = np.zeros((N, N))
# Loop through omega_M and omega_L and find the optimum likelihood.
- What is the optimum $\Omega_M$? Answer to 1 decimal place.
- What is the optimum $\Omega_{\Lambda}$? Answer to 1 decimal place.